Using Compression to Compare Objects
In my previous blog post, I discussed our endeavor to benefit from unsupervised learning on CyberPoint's malware dataset. One of the more intriguing tools I played with during that effort was the normalized compression distance (NCD).
NCD1 was introduced in 2004 as a universal distance function, with two very promising features:
- It's applicable to any sort of data you might want to analyze — without requiring any knowledge of the particulars of the data format or other domain knowledge.
- It approximately subsumes any other (computable) distance function you might come up with — even if it was cleverly devised with subject matter expertise on the particular type of data.
It achieves this by approximating the normalized Kolmogorov distance.
The Kolmogorov distance between two objects is actually pretty easy to conceptualize — it is the length of the shortest program that can transform one object into the other. Unlike many popular similarity measures, this provides a universal notion of similarity by quantifying the difference between two objects without restricting the type of difference. Unfortunately, this quantity is uncomputable, i.e., it is impossible to write a computer program that will compute it. So, Kolmogorov distance makes for an interesting philosophical discussion, but doesn't help us in our efforts to classify malware. Fortunately, the creators of NCD were able to use compression to approximate Kolmogorov distance, while (at least in theory) maintaining the desired universality properties. The idea is that, if you concatenate two similar objects and apply a compression algorithm, you're going to see a significant amount of compression.
Specifically, if we let C(X) note the result of compressing X with some compression algorithm C, and let |X| be the size of X, we have:
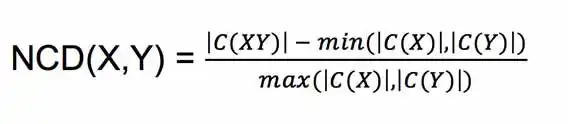
In words, NCD(X, Y) measures the mutual compressibility of X and Y, relative to the compressibility of X and Y individually.
The elegance of this appealed to me immensely, and I eagerly took to experimenting with clustering malware samples using NCD as the distance function.
When Compression Fails to Detect Similarity
As was the case in most of my unsupervised learning endeavors, I was thoroughly disappointed. While a few malware samples were deemed similar, most were determined to be equally dissimilar, yielding a couple of very small clusters, and a huge number of singletons. (While we weren't wowed with clustering using other distance functions, this was particularly egregious.) After initial experiments with bz2, I also tried lzma and zlib.
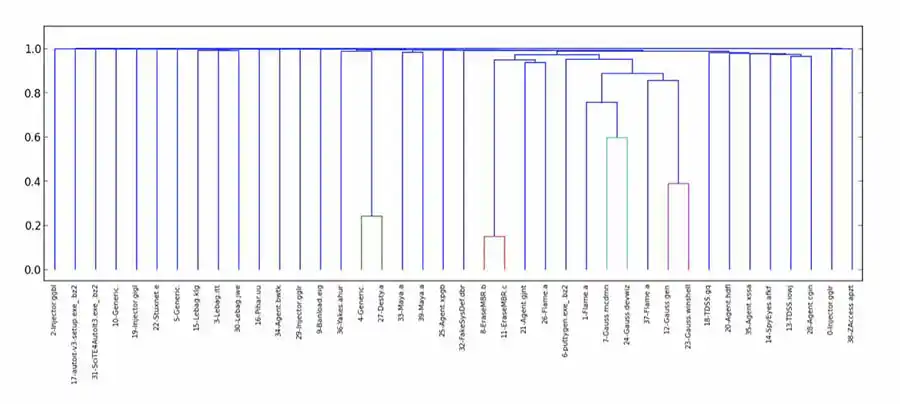
As I explored what went wrong, I found that NCD deemed many malware samples maximally dissimilar to themselves. That is, the distance from a sample to itself, which should have been 0, was actually 1 or higher — where, in theory 0 ≤ NCD(x, y) ≤ 1, for any samples x and y.
So, it turns out that there is a requirement that the compression algorithm be normal in order for NCD to satisfy the desired properties. A normal compression algorithm is one that satisfies (up to an additive O(log n) term) the following properties:
- Idempotency: C(xx) = C(x). (The algorithm takes sufficient advantage of the most simple redundancy in the string being compressed.)
- Monotonicity: C(xy) ≥ C(x). (If you add more data at the end of a string, it doesn't magically become significantly more compressible.)
- Symmetry: C(xy) = C(yx). (Swapping the order of the data doesn't make a significant impact on its compressibility.)
- Distributivity: C(xy)+C(z) ≤ C(xz)+C(yz). (This condition is less intuitive, but is roughly analogous to the triangle inequality.)
where n is the maximal binary length of any of the x, y, or z appearing in the statement.
The O(log n) leeway factor is somewhat perplexing — it refers to asymptotic behavior, but, in reality, we don't have objects with sizes approaching infinity. It's hard to see why the asymptotic complexity class would impact the outcome at any finite size.
Nonetheless, it seems that this is the problem. Unlike Kolmogorov complexity, compression algorithms are limited. The major hurdle is to effectively use information from string X for the compression of string Y in computing C(XY). Algorithms like bz2 and zlib have an explicit block size as a limiting factor; if |X| > block_size, then there is no hope of benefiting from any similarity between X and Y. In contrast, lzma doesn't have a block size limitation, but instead has a finite dictionary size; as it processes its input, the dictionary grows. Once the dictionary is full, it is erased and the algorithm starts with an empty dictionary at whatever point it has reached in its input. Again, if this occurs before reaching the start of Y, hope of detecting any similarity between X and Y is lost. Likewise, even if X is small, but Y is large, with the portion of Y that is similar to X appearing well into Y, the similarity can't be detected. This clearly poses a problem for idempotence and symmetry.2
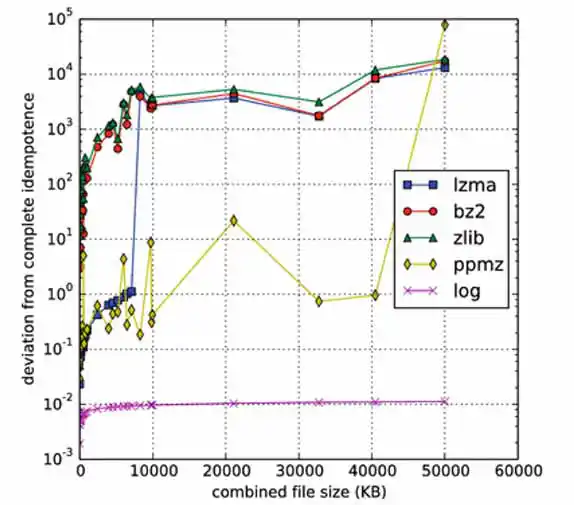
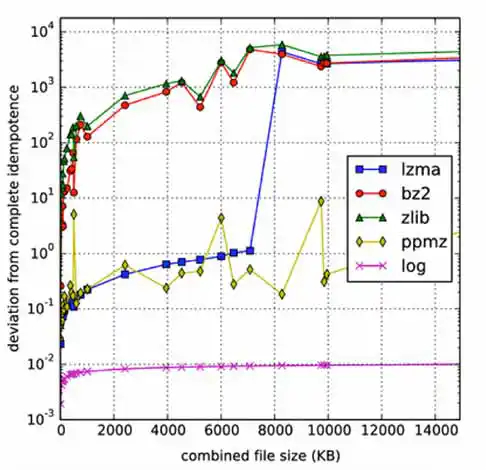
Improving NCD for Large Malware
In my recent paper in the Journal of Computer Virology and Hacking Techniques, I discuss these issues and explore an alternate definition of NCD, designed to handle large objects, in spite of the limitations of compression algorithms. Instead of concatenating the two objects to be compared, one combines them in some other way, with the goal of getting like parts of the two objects near each other, so that the compression algorithm can do its job. This showed much promise, as shown in Figure 4, where we measured the effectiveness of the different variants of NCD by using them with a k-nearest neighbor classifier to classify Android malware by malware family (with k=1 and k=5).
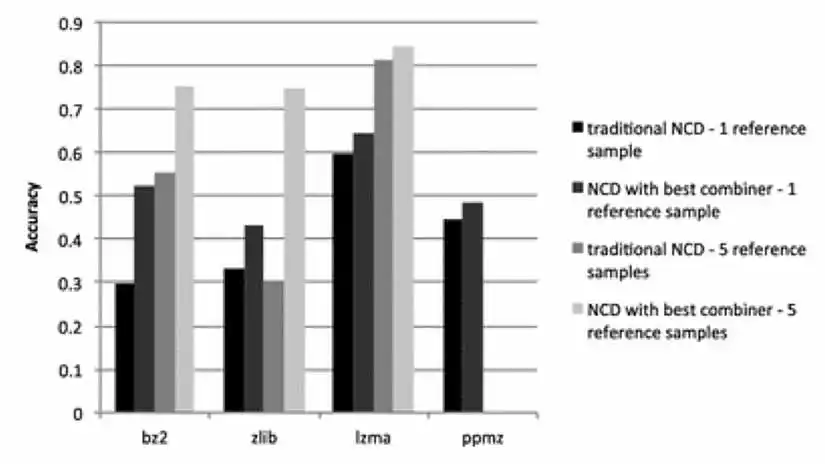
Note that Figure 4 shows the results for the best of several methods I tried for combining files.
Although these alternate combiners improved the usability of NCD for classification of large malware, it seems that experimentation would generally be necessary to determine which combiner/parameters are best for a particular dataset and compression algorithm. So, this could prove quite beneficial in cases where one has labeled data that can be used for such experiments (as I did). However, in the absence of labeled data, it leaves one, as usual, experimenting in the dark. Still, I remain hopeful that NCD can be adapted to fulfill its potential as a universal distance measure, even in the case of large, unlabeled, data.